Development of a cavitation enhancement technology to access archived tissues for epigenetic-based biomedical research
Dr. Samantha Pattenden, Associate Professor within the Division of Chemical Biology and Medicinal Chemistry, and Director of Applied Epigenetic Screening Technologies in the Center for Integrative Chemical Biology and Drug Discovery, and Dr. Ian Davis, Professor, Pediatrics and Genetics, and Associate Division Chief, Hematology/Oncology, UNC Lineberger were awarded a $1.6 million dollar grant from the National Institutes of General Medical Sciences (NIGMS). They will apply a cavitation enhancement technology developed in the laboratory of Dr. Paul Dayton in the Joint Department of Biomedical Engineering UNC and NC State to extract chromatin from archived tissues. Part of this work will be conducted in collaboration with Dr. Fei Zou and Dr. Di Wu in the Department of Biostatistics at UNC.
Innovative new technologies that enable experimentally robust interrogation of epigenetic mechanisms are needed to broaden our understanding of epigenetic regulatory pathways in human development, disease, and therapeutic resistance. Formalin fixed, paraffin embedded (FFPE) tissues contain a wealth of information on human disease, however, extraction of high-quality chromatin (DNA together with associated nuclear proteins) from these samples for use in epigenetic assays has proven virtually impossible. We will explore the use of a unique cavitation enhancement reagent in simplifying and standardizing chromatin extraction from FFPE tissues, with the goal of making archived biospecimens available for a broad range of epigenetic-based biomedical research.
Wang and Graham Awarded a 3-year $600,000 Eshelman Institute for Innovation Award to Develop Novel TYRO3 Degraders for the Treatment of Cancer and
Dr. Xiaodong Wang Receives R01 to Develop Novel TYRO3 Inhibitors for Treatment of Cancer
The TAM (TYRO3, AXL, MERTK) family of receptor tyrosine kinases play important roles in tumor survival by ectopic expression and/or immune suppression in the tumor microenvironment. While TAM kinase inhibitors have been therapeutically effective in preclinical cancer models, a growing body of evidence suggests that TAM kinases can also have kinase-independent functions. To disrupt these non-canonical functions, we will develop proteolysis-targeting chimera (PROTAC) degraders that selectively target TYRO3 for ubiquitination and degradation. The lead compounds will be tested in TYRO3-dependent immune-competent murine cancer models.
May 14, 2021
Dr. Xiaodong Wang, a Research Professor within the Center for Integrative Chemical Biology and Drug Discovery, and its Director of Medicinal Chemistry was awarded over $1.5 million from NIH along with her collaborator, Douglas Graham, PhD of Emory University. Both Wang and Graham have been working along with Dr. Shelley Earp of Lineberger Comprehensive Cancer Center and Dr. Stephen Frye of the CICBDD. This grant will support them to develop novel, potent, and selective TYRO3 inhibitors. TYRO3 is a member of the TAM (TYRO3, AXL, MERTK) family of receptor tyrosine kinases and is a potential therapeutic target in a wide variety of human tumors. Here, they propose to develop novel, potent, and selective TYRO3 inhibitors and validate their biochemical and functional activities in TYRO3-dependent tumor xenograft models and immune-competent syngeneic cancer models.
Stephen Frye Receives Maximizing Investigators’ Research Award
Tops $2 million, supports small molecule research to identify new approaches to modulating drug target
Brittany Jennings
March 26, 2021
Each day, Stephen Frye, Ph.D., works to identify pharmacologic approaches to intervene in novel drug targets within the human body with the goal of creating new therapeutic interventions that will directly aid patients.
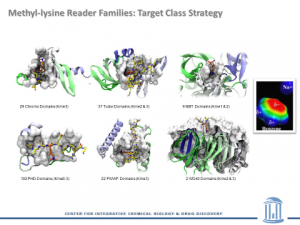
Today, he’s pushing his work further with the support of a prestigious Maximizing Investigators’ Research Award, a mechanism that aims to enhance investigator scientific productivity and the chances for important breakthroughs. The five-year grant totals $2,129,195 and will support his research project, “Probing Allostery In Methyl-Lysine Reader Domains.”
“This award will enable a broad, thematic approach to establishing allosteric interventions in an emerging class of drug targets, methyl-lysine reader domains,” said Frye, a Fred Eshelman Distinguished Professor and Co-Director of the Center for Integrative Chemical Biology and Drug Discovery. Find More about the Frye Grant Here
Wang receives $2.5 million grant to further novel cancer treatment research
Veronica Correa
April 9, 2020

Xiaodong Wang, Ph.D., has received a selective $2.5 million R01 grant from the National Institutes of Health to research and develop a new treatment for targeting tumor cells.
Wang is a research associate professor at the UNC Eshelman School of Pharmacy’s Center for Integrative Chemical Biology and Drug Discovery (CICBDD). Her lab focuses on developing new drugs or drug leads for new molecular targets. A MERTK/FLT3 dual inhibitor MRX-2843 from her lab is in phase I clinical trial for patients with relapsed or refractory solid tumors.
The five-year grant will investigate cancer treatments for the dual-inhibitor targets MERTK and AXL. Douglas Graham, MD, PhD from Emory University
Douglas Graham, MD, PhD from Emory University [and Dmitri Kireev, PhD from CICBDD are also PI’s.]
These two are regularly overexpressed in tumor cells, particularly lung cancer. Because both targets also have a role in suppressing an anti-tumor immune response and then contribute to further tumor growth, a dual-inhibition strategy would be more effective by providing anti-tumor action through both direct effects on cancer cells and modulation of the innate immune response, said Wang.
There are several small molecules in development that target MERTK and/or AXL. Wang said the lab has already identified an in vivo tool compound for research use. She plans to use a new computational method from the Kireev lab to develop and validate MERTK and AXL inhibitors.
Wang said that their research can have potential clinical applications, and even lead to the development of additional new drugs. She said it was exciting to get to conduct new work in this research field to be able to help patients.
“We really hope that we’re going to come out with a new preclinical candidate which will be ready to enter clinical trial after Investigational New Drug Application (IND) enabling,” she said.
New NIH proposal aims to advance Alzheimer’s drug discovery research

A group of researchers from the University of North Carolina at Chapel Hill are part of a new research proposal to create chemical and biological tools to explore novel molecular targets related to Alzheimer’s disease.
The proposal, which was submitted through the NIH National Institute on Aging, aims to create new chemical probes to validate therapeutic targets for Alzheimer’s, which has no known cure, and very few treatments. The project was awarded $37.5 million for five years. Emory University is the leader on the proposal, which is a collaboration between five institutions including UNC. The grant will establish the Open-AD Drug Discovery Center to advance Alzheimer’s treatment. Open-AD will develop a suite of target enabling tools including high quality antibodies and chemical probes, and openly disseminate all data, methods and reagents to any interested academic and/or commercial investigator to accelerate validation of novel drug targets and to seed new drug discovery efforts.
The UNC team, which is focused on medicinal chemistry, includes Stephen Frye, Ph.D., Director of the Center for Integrative Chemical Biology and Drug Discovery (CICBDD), and research assistant professors Alison Axtman, Ph.D. and Kevin Frankowski, Ph.D. Frye will be the Principal Investigator on the UNC end, while Frankowski and Axtman will be co-investigators in medicinal chemistry. Other UNC contributors include:
- Jeff Aubé, Ph.D., Co-Investigator, Medicinal Chemistry
- Dmitri Kireev, Ph.D., Co-Investigator, Computational Chemistry
- Xiaodong Wang, Ph.D., Co-Investigator, Medicinal Chemistry
- Kenneth Pearce, Ph.D., Co-Investigator, Lead Discovery, Characterization and Project Manager
- Brian Hardy, Compound and Laboratory Manager, Lead Discovery and Characterization
- Mike Stashko, B.S., Research Associate, Lead Discovery and Characterization
- Tim Willson, Ph.D., Co-Investigator, Medicinal Chemistry
- Carrow Wells, B.S., Research Associate, Medicinal Chemistry
Other participating institutions include the Structural Genomics Consortium (SGC) at the University of Oxford, Stanford University, and Sage Bionetworks in Seattle. Each group will focus on a specific part of the drug/probe discovery process, such as structural biology, bioinformatics, or assay development and high-throughput screening. The initiation of this NIH project is also contributing to a key UNC objective to increase its research involvement in neuroscience.
Frankowski said the project was an opportunity to work with renowned experts in Alzheimer’s research who have contributed to new theories of disease etiology and potential intervention points. A key challenge he mentioned in progressing this research is finding chemical probes that can penetrate the blood-brain barrier to reach targets in the central nervous system. The National Institute of Aging has previously funded the Accelerating Medicines Partnership-Alzheimer’s Disease (AMP-AD), which has provided more than 500 new candidate targets for Alzheimer’s disease, and these targets serve as the foundation for Open-AD focused on pharmacologic validation and drug discovery.
“However, despite years of research and looking into it, we don’t understand the fundamental causes for Alzheimer’s disease,” Frankowski said. The UNC effort will test disease hypotheses with chemical probes which can confirm or reject novel targets’ role in AD, and if confirmed, form the basis for discovery of new drugs.
The project will also follow an open drug discovery methodology. Axtman said lab notebooks and chemical probes will be released so others can access new discoveries. Following this approach, Open-AD will test the overarching hypothesis that open drug discovery will accelerate the development of AD medicines by catalyzing robust experimental evaluation of a series of complementary therapeutic hypotheses.
Pinnacle Hill Announces First Project Agreement
Brittany Jennings
December 18, 2019
Alliance Between UNC-Chapel Hill and Deerfield intends to advance select Carolina innovations toward a possible new treatment against multiple myeloma
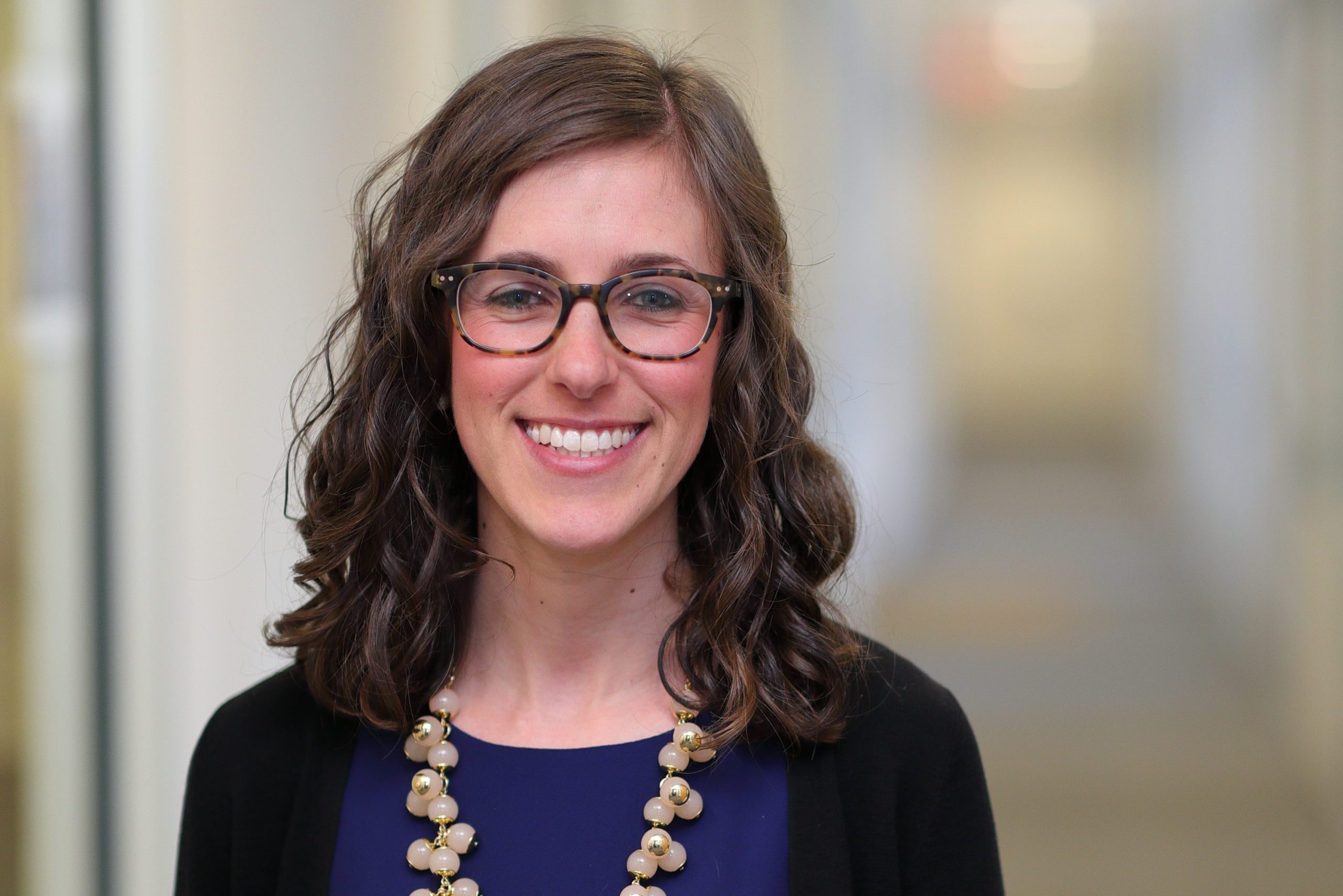

Pinnacle Hill, the research and development partnership between the University of North Carolina at Chapel Hill and Deerfield Management Company, an investment management firm, recently announced its first project agreement to advance the preclinical development of new medicines.
Pinnacle Hill will support a project of Lindsey James, an assistant professor in the UNC Eshelman School of Pharmacy’s Division of Chemical Biology and Medicinal Chemistry. Her work focuses on multiple myeloma, a devastating cancer that develops in bone marrow.
James and her research team are interested in epigenetic abnormalities that lead to cancer and are working to apply medicinal chemistry, chemical biology, and cancer biology approaches to discover potential therapeutic agents. Through this first project, Pinnacle Hill will devote targeted resources to advancing discoveries which have great therapeutic potential for patients with the disease.
“There’s no doubt this initial agreement from Pinnacle Hill will accelerate Dr. James’ life changing work,” said Vice Chancellor for Research and Pinnacle Hill Steering Committee Member Terry Magnuson. “The project is a testament to the power of the partnership between Deerfield and UNC-Chapel Hill.”
Launched in October 2018, Pinnacle Hill was established with a $65 million funding commitment from Deerfield to accelerate the development and translation of these biomedical and life-science innovations to address the significant unmet medical needs.
Pinnacle Hill focuses on drug research projects that are approved and overseen by a joint steering committee comprised of members from UNC-Chapel Hill and Deerfield leadership teams. Each selected project has the potential to receive funding to support investigational new drug enabling studies. UNC members of the joint committee are:
- Terry Magnuson, UNC-Chapel Hill’s vice chancellor for research and Kay M. & Van L. Weatherspoon Eminent Distinguished Professor of genetics
- Dr. Blossom Damania, the Boshamer Distinguished professor of microbiology and immunology and vice dean for research in the UNC School of Medicine
- John Bamforth, director, Eshelman Institute for Innovation
“The goal of our alliance with UNC-Chapel Hill is to identify and advance the most important discoveries that could improve the lives of patients and their families,” said James E. Flynn, Managing Partner at Deerfield. “Professor James’ work exemplifies this and is outstanding. We are delighted to advance her novel research and are optimistic about continued progress in Dr. James’ laboratory as well as other milestones to come at UNC. They have been a great partner.”
Eshelman Institute for Innovation Awards Students, Postdocs
Brittany Jennings
December 20, 2019
The Eshelman Institute for Innovation (EII) recently completed their 4th EII Student/Postdoc Project award cycle, awarding $25,000 to three student teams totaling $75,000. Lead Innovators and project titles include:
Bryce Hart, Development of a small molecule ligand for DCAF1 (PhD student in Chemical Biology and Medicinal Chemistry Division) 
Sabrina Iskander, Exploiting Promiscuous tRNA Synthetases to Identify Non-Canonical Peptide Inhibitors of KRAS via mRNA Display (PhD student in Center for Integrative Chemical Biology and Drug Discovery Center)
Jasmine King, In-situ Thermogelling injectable hydrogel to enhance cytotoxic neural stem cell delivery for post-surgical Glioblastoma multiforme (PhD student in Pharmacoengineering and Molecular Pharmaceutics Division)
Please join the EII in congratulating their selected participants and we look forward to seeing what the students will accomplish! The EII would also like to thank their expert reviewers for their support during this grant cycle.
To learn more about the EII, visit our website or email eshelmaninstitute@unc.edu.








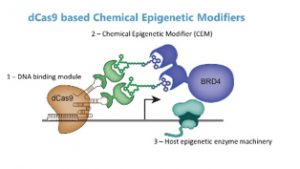
 Nathaniel Hathaway, Ph.D., a researcher at the UNC Eshelman School of Pharmacy’s Division of Chemical Biology and Medicinal Chemistry, Center for Integrative Chemical Biology and Drug Discovery and the Lineberger Comprehensive Cancer Center, was published in Nature Biotechnology earlier this month for his work creating tools that can control gene expression.Hathaway, who will be promoted to associate professor Dec. 1, focuses on understanding the mechanisms of gene regulation by chromatin while discovering small molecules that change epigenetic pathways. The Nature Biotechnology publication was a collaboration with other researchers from UNC, a group at Mount Sinai Center for Therapeutics Discovery lead by Dr. Jian Jin, and the Khan research group at National Institutes of Health’s National Cancer Institute.This latest research, titled “
Nathaniel Hathaway, Ph.D., a researcher at the UNC Eshelman School of Pharmacy’s Division of Chemical Biology and Medicinal Chemistry, Center for Integrative Chemical Biology and Drug Discovery and the Lineberger Comprehensive Cancer Center, was published in Nature Biotechnology earlier this month for his work creating tools that can control gene expression.Hathaway, who will be promoted to associate professor Dec. 1, focuses on understanding the mechanisms of gene regulation by chromatin while discovering small molecules that change epigenetic pathways. The Nature Biotechnology publication was a collaboration with other researchers from UNC, a group at Mount Sinai Center for Therapeutics Discovery lead by Dr. Jian Jin, and the Khan research group at National Institutes of Health’s National Cancer Institute.This latest research, titled “